Hi everyone, I am new to this community - first and foremost, thank you for having this tool available to the public and providing support!
I am working on a sleep dataset: EDF and XML files, and have couple questions:
1) How to resolve: "Fatal error - Could not load: <C:.....XML>" while loading my XML file.
2) All PSG signals are displayed in "mV" units - Is there any way to convert mV to actual physiological units, for example: SpO2 readings to % ?
3) The code runs very slow on my computer - Any recommendations on minimum hardware requirements?
Thank you in advance! Best, Onur
Hi Onur,
It's great to hear from you and we are happy to help.
For your first question, it seems you tried to load your own XML annotation, but EDF Viewer is created to work with NSRR provided annotation files.
Secondly, EDF Viewer does visualize signals with actual physiological units. For example, we show pulse with unit BPM and SpO2 with % (see images below).
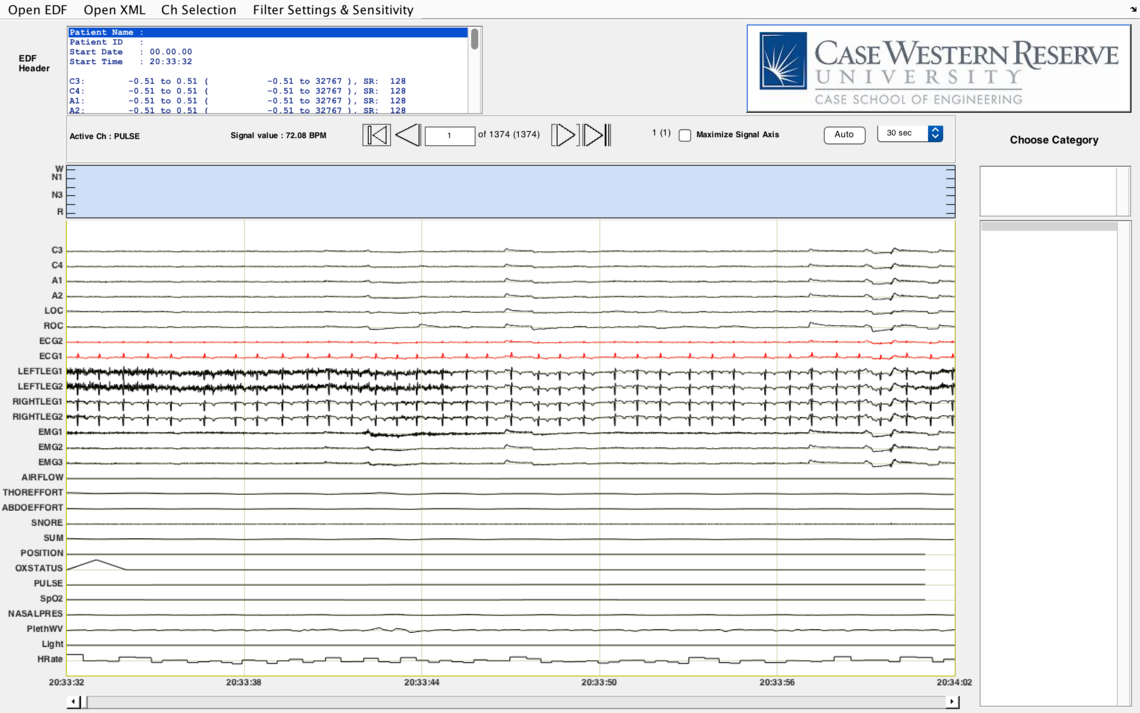
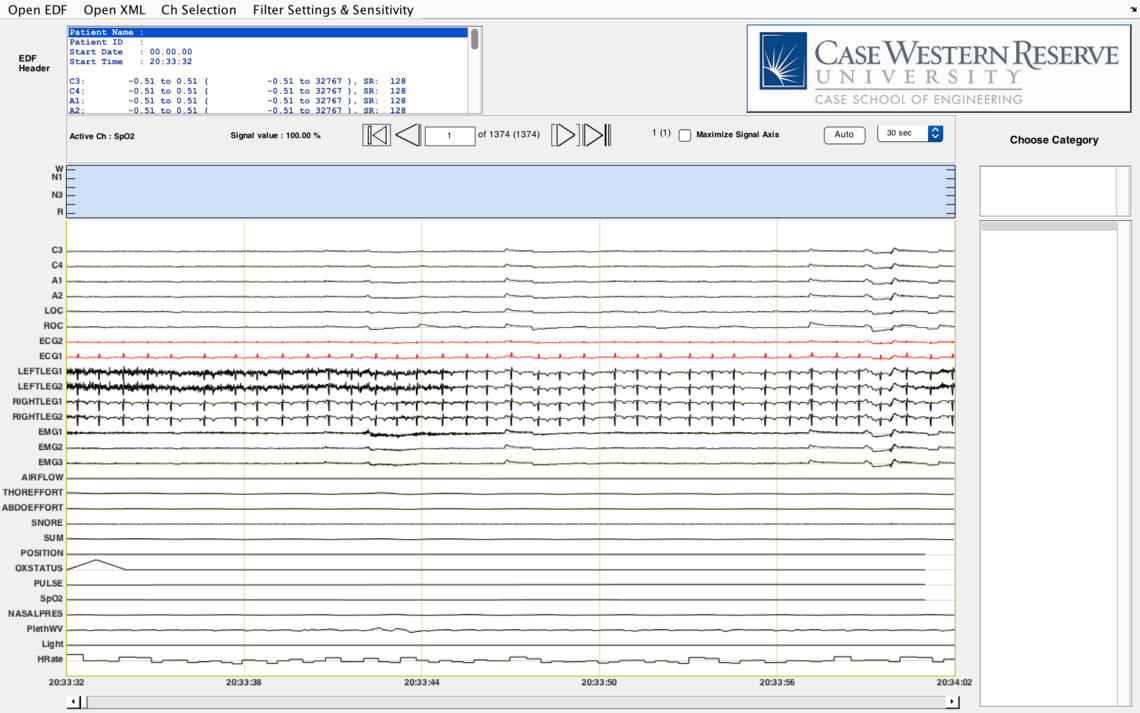
Responding to your last question, the loading of EDF and annotations takes a couple of minutes and it may take longer depending on the size of files. Our EDF Viewer requires MATLAB Compiler Runtime version 8.5 (MATLAB R2015a) to run, which needs operating system version to be >= Windows XP or >= Mavericks if you are a MAC user. We also recommend at least 2GB ram to ensure the application runs properly.
We can better help with your problems if you can provide more information about what EDF and annotation you tried to load and what operating system you were working on.
Thanks!